| 1 |
dpavlin |
1.1 |
<p><b>Scientific Background</b></p> |
| 2 |
|
|
<p>Mismatch Repair is a fundamental process, found in all living cells from bacteria |
| 3 |
|
|
to humans, and functions to maintain the integrity of DNA and the separation |
| 4 |
|
|
of species. Mismatch Repair Systems functions to:</p> |
| 5 |
|
|
<ul> |
| 6 |
|
|
<li>repair errors made during DNA replication</li> |
| 7 |
|
|
<li>provide a genetic barrier to speciation by preventing genetic recombination |
| 8 |
|
|
between totally or partially divergent species.</li> |
| 9 |
|
|
</ul> |
| 10 |
|
|
<p>Mutations in or inactivation of the Mismatch Repair genes allow high levels |
| 11 |
|
|
of recombination between divergent DNA sequences, including genomes of different |
| 12 |
|
|
species. Normally, recombination between regions of such homeologous (partially |
| 13 |
|
|
homologous) DNA occurs at a negligible rates. However, when Mismatch Repair |
| 14 |
|
|
is not operating, recombination frequencies increase by more than 1000 fold |
| 15 |
|
|
(see figure).</p> |
| 16 |
|
|
<p>The role of Mismatch Repair in recombination and speciation was discovered |
| 17 |
|
|
by Prof. Miroslav Radman and his co-workers, a world renown group in the fields |
| 18 |
|
|
of DNA repair and "Homeologous Recombination". The patents describing the broad |
| 19 |
|
|
application of this discovery were the basis for founding MIXIS France.</p> |
| 20 |
|
|
<p><b>Technology</b></p> |
| 21 |
|
|
<p>MIXIS technology mimics the molecular strategy of the immune system to generate |
| 22 |
|
|
diversity. Inhibition of Mismatch Repair allows recombination between homeologous |
| 23 |
|
|
sequences to generate mosaics of pre-existing sequences, and elevates mutation |
| 24 |
|
|
rates to create de novo variation.</p> |
| 25 |
|
|
<p>MIXIS core technology allows single genes or even genomes of interest to be |
| 26 |
|
|
driven through an accelerated evolution process (see figure): Homeologous parental |
| 27 |
|
|
genes or genomes can be recombined repeatedly under Mismatch Repair deficient |
| 28 |
|
|
conditions, resulting in mosaic genes and genomes. These novel genes will code |
| 29 |
|
|
for novel proteins or biosynthesis products.</p> |
| 30 |
|
|
<p>With an appropriate screening system, the MIXIS technology can allow identification |
| 31 |
|
|
of gene products - low molecular weight entities, proteins, organisms - with |
| 32 |
|
|
desired properties.</p> |
| 33 |
|
|
<p>Because of its <b>simplicity</b>, <b>speed</b> and <b>versatility</b> MIXIS |
| 34 |
|
|
technology represents a unique and powerful tool in the compound discovery and |
| 35 |
|
|
development process.</p> |
| 36 |
|
|
<p align="center"><img src="p/mixisGene.gif" width="270" height="379"></p> |
| 37 |
|
|
<p>The MIXIS method is easy to use (in vivo recombination) and is, therefore, |
| 38 |
|
|
quantitatively and qualitatively the most powerful method available for generation |
| 39 |
|
|
of repertoires of genetic diversity.</p> |
| 40 |
|
|
<p>A critical advantage of the MIXIS technology is its ability to produce diversity |
| 41 |
|
|
within complex metabolic pathways without requiring a detailed knowledge of |
| 42 |
|
|
the genes involved.</p> |
| 43 |
|
|
<p>Thanks to a strategic alliance with PLIVA Research Institute in Zagreb, Croatia, |
| 44 |
|
|
MIXIS has access to all upstream activities necessary to screen, identify, characterise |
| 45 |
|
|
and produce novel recombinant genes and their products.</p> |
| 46 |
|
|
<p><b>Applications</b></p> |
| 47 |
|
|
<p>MIXIS technology can be applied to several fields of life science research |
| 48 |
|
|
and development:</p> |
| 49 |
|
|
<ul> |
| 50 |
|
|
<li>generation of novel, low molecular weight chemical entities, even those |
| 51 |
|
|
produced by complex metabolic pathways, for use as pharmaceuticals, crop protection, |
| 52 |
|
|
etc.</li> |
| 53 |
|
|
<li>optimisation of activities of macromolecules such as proteins, (substrate |
| 54 |
|
|
specificity, stability, activity, etc.)</li> |
| 55 |
|
|
<li>generation of organisms with desired properties, such as micro-organisms |
| 56 |
|
|
designed for enhanced production under stressful conditions or with modified |
| 57 |
|
|
substrate specificity, as might be required for environmental protection applications. |
| 58 |
|
|
</li> |
| 59 |
|
|
</ul> |

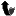 Parent Directory
|
Parent Directory
|  Revision Log
Revision Log